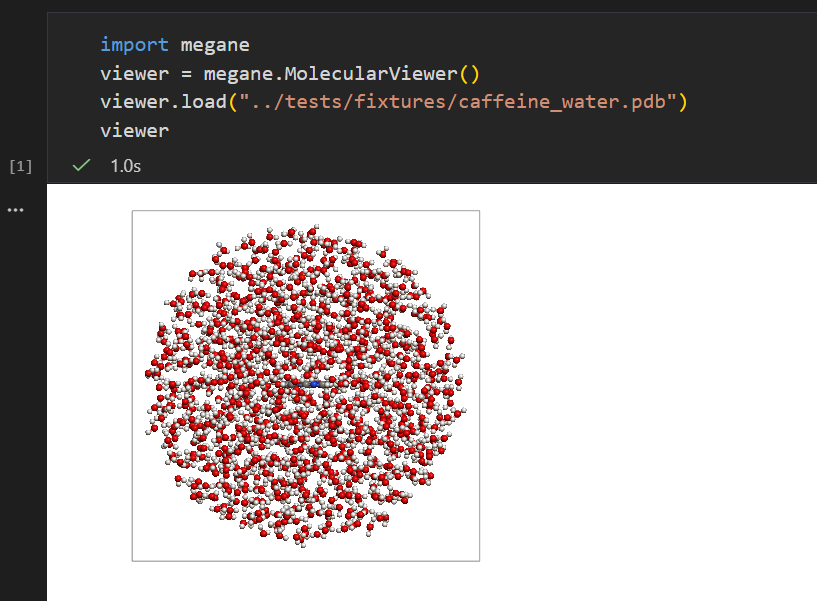
🚀 1M+ Atoms at 60fps
Billboard impostor rendering scales from small molecules to massive protein complexes in real time. Stream XTC trajectories over WebSocket — scrub thousands of frames without loading everything into memory.
🌍 Runs Everywhere
Jupyter widget, CLI server, React component, VSCode extension. Rust parsers (PDB, GRO, XYZ, MOL, CIF, XTC, LAMMPS, .traj) shared between Python (PyO3) and browser (WASM): parse once, run anywhere.
🧩 Visual Pipeline Editor
Build visualization workflows by wiring 11 node types — load data, filter atoms, adjust styles, generate labels, render coordination polyhedra, overlay vectors. No code required. 7 typed data channels flow through color-coded edges. An AI generator can build pipelines from natural language.
🔗 Embed & Integrate
Control the viewer from Plotly via ipywidgets events. Embed in MDX / Next.js docs. React to frame_change, selection_change, and measurement events. Use the framework-agnostic renderer from Vue, Svelte, or vanilla JS.
Start in your environment
megane works everywhere — pick your entry point.
Python / Jupyter
pip install meganeInteractive widget inside Jupyter notebooks. Build pipelines in Python, display structures inline.
Jupyter Guide →TypeScript / React
npm install megane-viewerDrop <PipelineViewer /> into any React app. Build pipelines with the TypeScript builder API.
React Guide →Docker
docker run hodakamori/meganeServe local structure files and view them instantly in the browser. No code needed.
CLI Guide →VSCode
Install from MarketplaceOpen .pdb, .gro, .xyz, .mol, .cif files directly in VS Code with the megane extension.
VSCode Extension →Scale
megane renders over 1 million atoms at 60fps in the browser. Small systems get high-quality InstancedMesh spheres and cylinders; large systems automatically switch to GPU-accelerated billboard impostors. No desktop app, no plugin — just a browser tab.
Trajectory streaming works over WebSocket via a binary protocol. Load an XTC file and scrub through thousands of frames in real time, without reading everything into memory.
Anywhere
One codebase, every environment.
| Environment | How | Install |
|---|---|---|
| Jupyter | anywidget inline viewer | pip install megane |
| Browser | megane serve local server | pip install megane |
| React | <MeganeViewer /> component | npm install megane-viewer |
| VSCode | Custom editor for .pdb, .gro, .xyz, .mol, .sdf, .cif | Extension |
The secret: PDB, GRO, XYZ, MOL, CIF, XTC, LAMMPS, and ASE .traj parsers are written in Rust and compiled to both PyO3 (Python) and WASM (browser). Parse once, run anywhere.
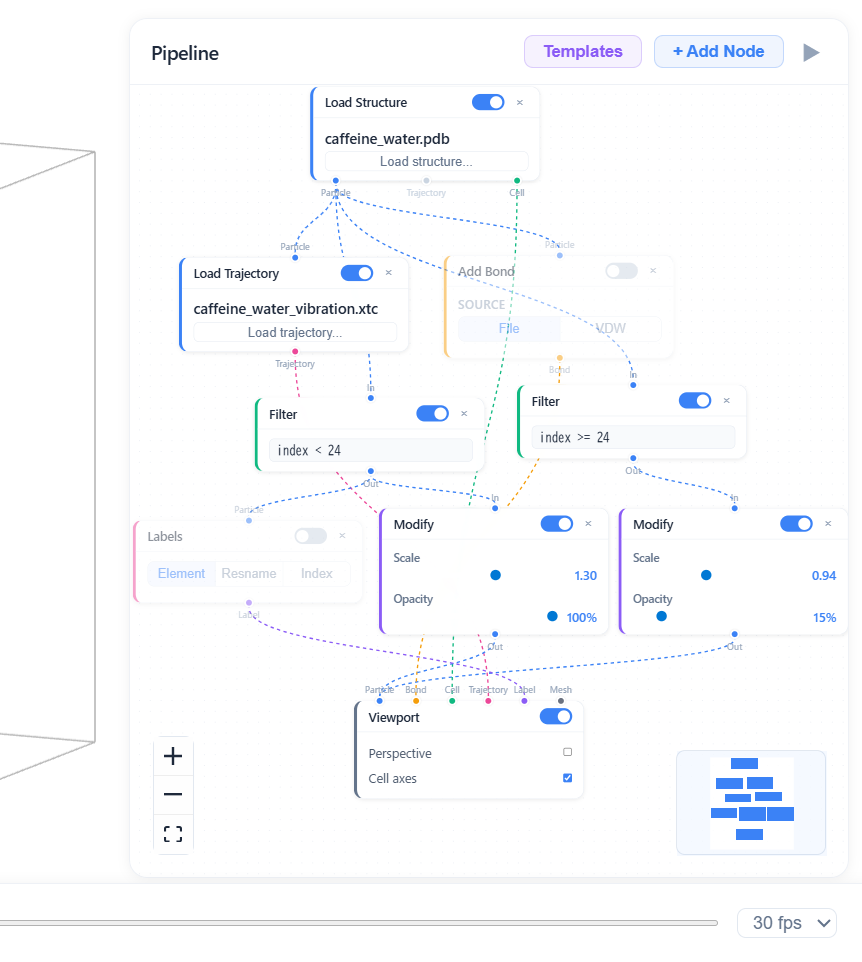
Visual Pipelines
Wire nodes to build visualization workflows — no code required.
11 node types across 5 categories: load data (structure, trajectory, streaming, vector), process (filter, modify), overlay (bonds, labels, polyhedra, vectors), and display in a 3D viewport.
7 typed data channels — particle, bond, cell, label, mesh, trajectory, vector — flow through color-coded edges. Only matching types can connect.
Pipelines serialize to JSON, so you can save, share, and version-control your visualization recipes.
Integrate
megane is not a walled garden. It fits into your existing workflow.
Plotly — Click a point on a Plotly FigureWidget to jump to a trajectory frame. Use megane's on_event("frame_change") callback to update Plotly markers in sync.
MDX / Next.js — Drop <MeganeViewer /> or <Viewport /> into your .mdx documentation. WASM parsing works out of the box with a one-line webpack config.
ipywidgets — React to frame_change, selection_change, and measurement events. Compose megane with any widget in the Jupyter ecosystem.
Framework-agnostic — MoleculeRenderer is a plain Three.js class. Mount it in Vue, Svelte, or a vanilla <div>.